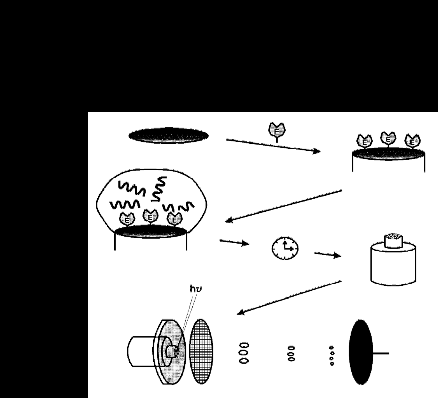
Once the Human Genome has been sequenced and entered into databases, it will be possible to search for codons corresponding to given amino acid sequences. Therefore, the ability to rapidly characterize proteins becomes an important experimental consideration. We have taken some preliminary steps in this direction by developing a rapid and sensitive method for the mass spectrometric characterization of peptides. A schematic of the process is shown below.

Probe tips are first prepared, in bulk, by covalent attachment of enzymes to the probe surface (in this case through DSP activation of a gold surface followed by enzyme linkage via a primary amine). The probe tips can then be used, at will, for protein characterization by application of the protein in an appropriate buffer and allowing time for digestion, which is usually performed at elevated temperature. Digestion is stopped with the addition of a MALDI matrix, and the reaction product/ matrix mixture allowed to dry (for the case of solid-phase matrices). The probe tips are then inserted directly into the mass spectrometer and MALDI-analyzed.
Shown below are the results of a 10 min. (40 oC) Au/trypsin/MALDI map of chicken egg lysozyme.
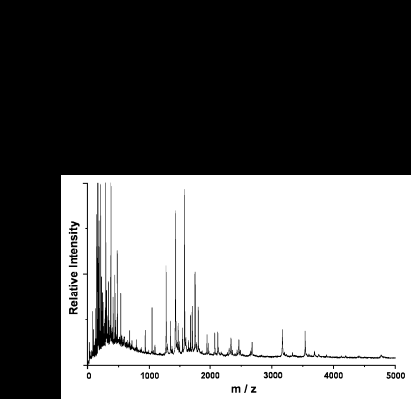
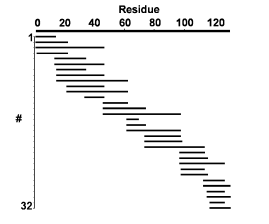
Lysozyme (10 pmol) was applied to the probe tip in the presence of the reducing agent DTT. 33 of a possible 67 tryptic fragments are observed in the m/z range of ~1000 - 6000 Da. The coverage map shows cleavage at 15 of the possible 17 sites and a three-fold mapping redundancy across the entire sequence.
The probe tips have also been used for partial sequence determination as demonstrated in the following spectrum.
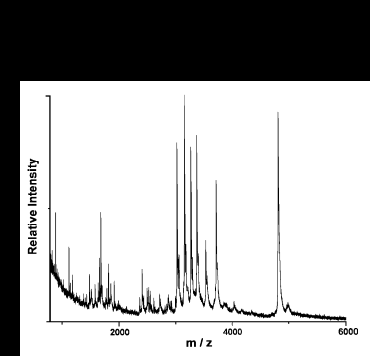
Myotoxin a (6 pmol) was digested for 15 min. at 45 oC using an Au/a-chymotrypsin probe tip, followed by addition of carboxypeptidase P (CPP). A set of ladder sequence peaks are observed, and from the mass difference between these peaks, the sequence -MDCRW- is determined. Using the parent molecular weight and the short piece of sequence, the OWL database was searched, and the identity of myotoxin was confirmed.
![[ <- ]](images/docsleft.gif) Back to Lab Page
Back to Lab Page