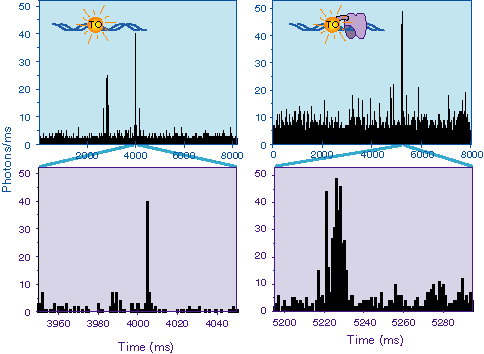
Part of a Study performed with the DNA intercalating dye,
thiazole orange (TO) is shown above. Here we compared the binding
of TO to DNA by itself or covalently attached to a zinc finger
DNA binding protein. In both cases, the TO only fluoresces when
bound to DNA. Thus one is watching the TO bind and leave the
DNA with time. One can see that the Zn finger-TO complex binds
more often and longer than does the free TO. However, on average,
the fluorescence intensity from the complex is less, probably
due to constrains placed on the TO by the covalent attachment.
1. JK Gimzewski & C Joachim (1999) Nanoscale science of
single molecules using local probes. Science 283: 1683-1688
2. AD Mehta, M Rief, JA Spudich, DA Smith & RM Simmons
(1999) Single-molecule biomechanics with optical methods. Science
283: 1689-1695
3. WE Moerner & M Orrit (1999) Illuminating single molecules
in condensed matter. Science 283: 1670-1676
4. S Weiss (1999) Fluorescence spectroscopy of single biomolecules.
Science 283: 1676-1683
5. SM Nie, DT Chiu & RN Zare (1994) Probing individual
molecules with confocal fluorescence microscopy. Science 266:
1018-1021
6. DC Daniel, M Thompson & NW Woodbury (1999) Fluorescence
intensity fluctuations of individual labeled DNA fragments and
a DNA binding protein in solution at the single molecule level:
A comparison of photobleaching, diffusion, and binding dynamics.
J. Phys. Chem. B (submitted):
7. L Lee, C-H Chen & L Chiu (1986) Thiazole orange - A
new dye for reticulocyte analysis. Cytometry 7: 508-517
8. M Thompson & NW Woodbury (1999) Fluorescent and photochemical
properties of a single zinc finger crosslinked with a fluorescent
DNA-binding probe. Biochemistry (submitted):
9. AA Deniz, M Dahan, JR Grunwell, T Ha, AE Faulhaber, DS
Chemla, S Weiss & PG Schultz (1999) Single-pair fluorescence
resonance energy transfer on freely diffusing molecules: Observation
of Förster distance dependence and subpopulations. Proc.
Natl. Acad. Sci. USA 96: 3670-3675
10. T Ha, TA Laurence, DS Chemla & S Weiss (1999) Polarization
spectroscopy of single fluorescent molecules. J. Phys. Chem.
B 103: 6839-6850
11. T Ha, AY Ting, J Liang, WB Caldwell, AA Deniz, DS Chemla,
PG Schultz & S Weiss (1999) Single-molecule fluorescence
spectroscopy of enzyme conformational dynamics and cleavage mechanism.
Proc. Natl. Acad. Sci. USA 96: 893-898
12. J Schaffer, A Volkmer, C Eggeling, V Subramaniam, G Striker
& CAM Seidel (1999) Identification of single molecules in
aqueous solution by time-resolved fluorescence anisotropy. J.
Phys. Chem. A 103: 331-336
13. S Wennmalm & R Rigler (1999) On death numbers and
survival times of single dye molecules. J. Phys. Chem. B 103:
2516-2519
14. RM Dickson, DJ Norris, Y-L Tzeng & WE Moerner (1996)
Three-dimensional imaging of single molecules solvated in pores
of poly(acrylamide) gels. Science 274: 966-968
15. AM Van Oijen, M Ketelaars, J Köhler, TJ Aartsma &
J Schmidt (1999) Unraveling the electronic structure of individual
photosynthetic pigment-protein complexes. Science 285: 400-402
16. AM Van Oijen, M Ketelaars, J Köhler, TJ Aartsma &
J Schmidt (1998) Spectroscopy of single light-harvesting complexes
from purple photosynthetic bacteria at 1.2 K. J. Phys. Chem.
B 102: 9363-9366
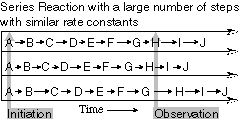 There are three major advantages
of looking at the dynamics and spectroscopy of single biological
molecules. The first is the study of processive or sequential
dynamics (Fig. 1). A population of molecules undergoing a series
of events, such as a complex assembly process or linear polymerization,
become out of phase and the detailed dynamics of individual steps
are lost. Of course, on the single molecule level, one watches
an individual molecule throughout the course of events, removing
the phase problem.
There are three major advantages
of looking at the dynamics and spectroscopy of single biological
molecules. The first is the study of processive or sequential
dynamics (Fig. 1). A population of molecules undergoing a series
of events, such as a complex assembly process or linear polymerization,
become out of phase and the detailed dynamics of individual steps
are lost. Of course, on the single molecule level, one watches
an individual molecule throughout the course of events, removing
the phase problem.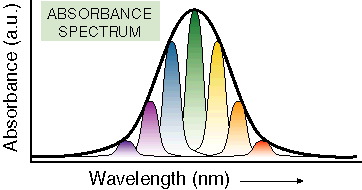 conformational
states with a particular solvent environment. Observing only population
averages can hide dynamic or mechanistic features of biological
molecules that are important to function. It can also have a profound
effect on the optical spectra of molecules, particularly at low
temperature where the solvent environment and the conformation
are not changing rapidly. Working with single molecules allows
one to measure optical spectra in the absence of inhomogeneous
broadening, learning both a great deal about the number and identity
of overlapping transitions as well as the underlying conformational
heterogeneity that gives rise to broad spectra in populations
of molecules (Fig. 2).
conformational
states with a particular solvent environment. Observing only population
averages can hide dynamic or mechanistic features of biological
molecules that are important to function. It can also have a profound
effect on the optical spectra of molecules, particularly at low
temperature where the solvent environment and the conformation
are not changing rapidly. Working with single molecules allows
one to measure optical spectra in the absence of inhomogeneous
broadening, learning both a great deal about the number and identity
of overlapping transitions as well as the underlying conformational
heterogeneity that gives rise to broad spectra in populations
of molecules (Fig. 2).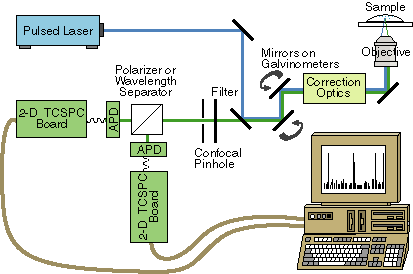 Normally, laser scanners are sold
as a unit with detectors for standard scanning confocal work.
Both the type of detector and the optics preceding it will have
to be modified for single molecule fluorescence work. After the
scanning mirrors, the fluorescence will be sent to a home built
confocal pinhole arrangement followed by either polarization or
wavelength selection optics, an avalanche photodiode (APD) detector
and a 2-dimensional timing board for recording time of arrival
and excited state lifetime, as shown in Fig. 5.
Normally, laser scanners are sold
as a unit with detectors for standard scanning confocal work.
Both the type of detector and the optics preceding it will have
to be modified for single molecule fluorescence work. After the
scanning mirrors, the fluorescence will be sent to a home built
confocal pinhole arrangement followed by either polarization or
wavelength selection optics, an avalanche photodiode (APD) detector
and a 2-dimensional timing board for recording time of arrival
and excited state lifetime, as shown in Fig. 5.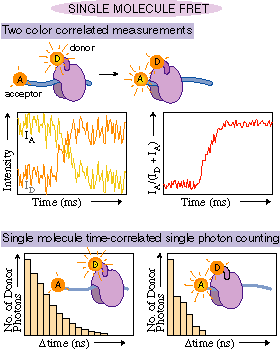 Correlation of two different wavelength
regions of emission is usually done when performing fluorescence
resonant energy transfer (FRET) experiments (e.g. ). Here, the
emission is again split into two detector channels, but this time
based on what wavelength range it is in. Consider the experiment
outlined in Fig. 6. As the two fluorophores approach each other,
the amount of energy transfer increases. As a result, the fraction
of the emission observed from the acceptor fluorophore increases
at the expense of the emission from the donor fluorophore. Again,
by correlating the changes in amplitude at two wavelength regions,
uncorrelated noise is suppressed.
Correlation of two different wavelength
regions of emission is usually done when performing fluorescence
resonant energy transfer (FRET) experiments (e.g. ). Here, the
emission is again split into two detector channels, but this time
based on what wavelength range it is in. Consider the experiment
outlined in Fig. 6. As the two fluorophores approach each other,
the amount of energy transfer increases. As a result, the fraction
of the emission observed from the acceptor fluorophore increases
at the expense of the emission from the donor fluorophore. Again,
by correlating the changes in amplitude at two wavelength regions,
uncorrelated noise is suppressed.